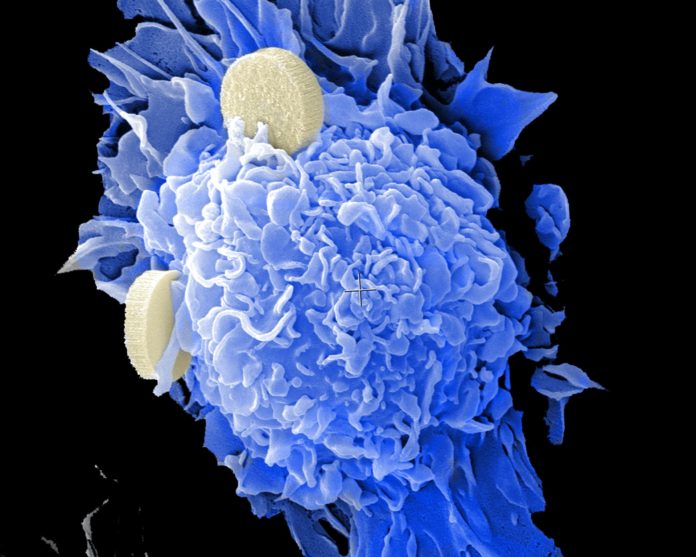
The Science and Engineering Research Board (SERB) has approved support for a study by the National Centre for Veterinary Type Cultures (NCVTC), ICAR-NRC from Hisar in Haryana, which will screen their library of 94 small molecule chemical inhibitors for antivirals against coronaviruses.
The molecules are known to inhibit cellular kinases, phosphatases, and epigenetic regulators such as histone methyltransferase, histone deacetylase, and DNA methyltransferase. The targets of these inhibitors are well characterized in cancer. However, their role in the virus life cycle is not known. The selected candidates (hits) with anti-coronavirus activity will be subjected to study their molecular mechanism of action, besides examining the generation of potential drug-resistant virus variants.
Classically, antiviral drugs are developed by directly targeting certain viral proteins. However, this strategy often fails due to the rapid generation of drug-resistant viruses. Unlike higher organisms, a viral polymerase-the viral enzyme that synthesizes its nucleic acid (RNA) does not have proofreading capacity. Therefore, RNA viruses such as the coronaviruses do not have the mechanisms to remove wrongly incorporated nucleotides (building blocks of viral RNA) during the synthesis of the viral genome. The lack of proofreading capacity results in the accumulation of point mutations in the viral genome. This leads to changes in viral proteins. The altered viral proteins may then become resistant to the available antiviral drugs. This intriguing ability of the viruses to rapidly and frequently change themselves is a big challenge for the scientists in developing antiviral drugs.
Viruses can only replicate inside the host cell. A host (human) cell contains around 25,000 proteins. During replication, viruses establish numerous interactions with these cellular proteins. A virus needs more than 1000 different cellular proteins to replicate inside the host cell effectively.
“Chemical library screening in medicinal chemistry research is a useful methodology that considerably shortens drug discovery and development cycle, especially for newly identified etiologic agents, such as SARS-Cov-2. Such an approach provides for rapid access to useful pharmacophores and narrows down the search for preferred molecular scaffolds and is also compatible with high-throughput robotic assays. While supporting Cov-2 vaccine development programs, it is also imperative to have sufficient focus toward anti-coronavirus drug discovery pipeline,” said Sandeep Verma, secretary, SERB.
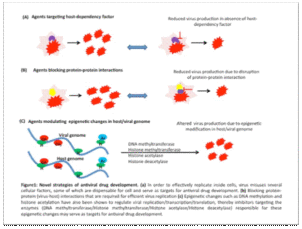
Naveen Kuman, scientist NCVTC is exploring an alternate strategy to target such cellular proteins, protein-protein (virus-host) interaction, or epigenetic regulators for antiviral drug development commonly called as host-directed antiviral therapy.
The host-directed antivirals are believed to have fewer tendencies in inducing drug resistance because it is not possible for the virus to change missing cellular functions by mutations easily. In addition, host-directed antiviral agents are likely to exert broad-spectrum antiviral effects because the requirement of host factors by viruses is usually conserved across the members of a particular virus family or sometimes even across the members of different virus families. The institute’s collection of small molecule inhibitors will be the resource from which Dr. Kumar hopes to find his antiviral weapon targeting cellular proteins, protein-protein (virus-host) interaction, or epigenetic regulators for Covid-19.